I get butterflies in my stomach when I hear ‘deoxyribose nucleic acid’!
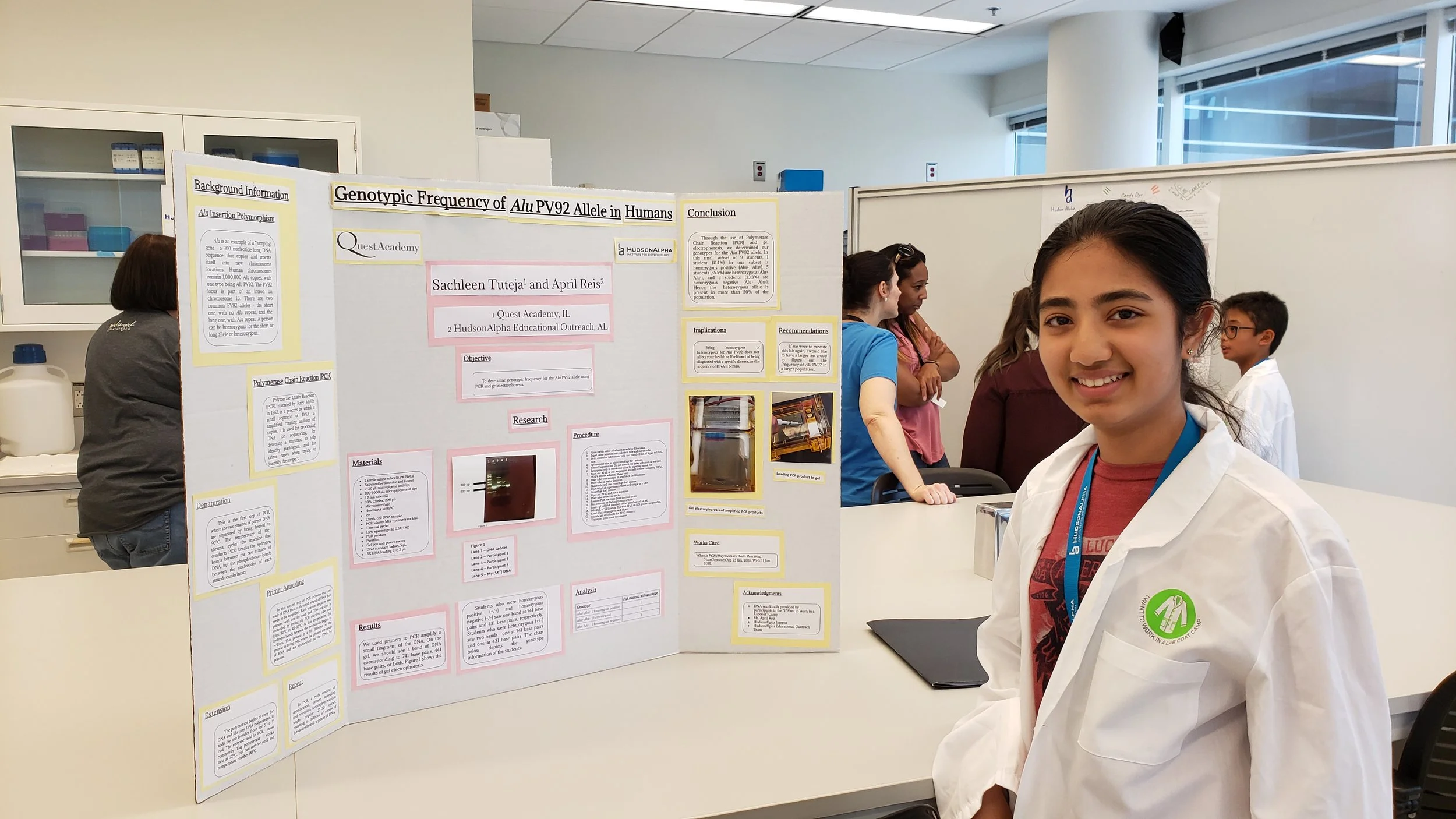
For years now, I have been in love in with DNA - the most powerful biological molecule on this planet! Ever since an introduction to this wonderous molecule in my elementary school science class while happily crushing strawberries, I’ve monkeyed around the rungs of DNA’s 3 billion base pair long double helical structure, jumping from one opportunity to another. Leveraging enrichment opportunities at selective summer camps at the Hudson Alpha Institute of Biotechnology and University of Notre Dame’s DNA learning center, passionately founding my school’s first Science Olympiad Team, educating about National DNA day, and excitedly sharing my scientific research at the Illinois Junior Academy of Science’s Science Fair - my love affair with DNA grew organically during middle school.
In high school, I found myself appreciating the nuances of DNA’s subtle variations that contribute to all life on this planet - by pursuing a research internship in molecular diagnostics under the guidance of Dr. Kai Lee Yap at Lurie Children’s Hospital of Chicago. Witnessing the harsh reality of DNA’s immense disruptive power to cause suffering and disease, my intellectual love affair with DNA transformed into a passionate pursuit of improved human health. I started pursuing a second internship in population health informatics with Dr. Theresa Walunas at Northwestern’s Center for Health Information Partnerships, enrolled in Pathophysiology and genomic data specialization courses, and started sharing my passion with the scientific community at large at various national and international professional conferences.
On an invigorating journey towards advancing human health!
Research Project - As the scientific community continues to discover novel genetic variants associated with human disorders, there is a growing need for a standardized approach for reporting these genetic variants, as accurate variant identification is crucial for effective diagnosis and treatment. In this study, using a manually curated set of 296 variants, I evaluated the performance of three variant annotation tools (Variant Effect Predictor, Alamut Batch, and ANNOVAR) for implementation in the clinical bioinformatics pipeline at the Molecular Diagnostics Laboratory at Lurie Children’s Hospital. This study is novel in its simultaneous assessment of multiple open source and licensed gene annotation software in the clinical setting towards a more accurate disease diagnosis.
Research Project - In this study, I am investigating the genetic relationship between risk scores of related autoimmune diseases, conditions in which the body’s natural defense systems mistakenly attack its own healthy cells and tissues. Treatment options for many autoimmune diseases are rather limited, and since a variety of other conditions mimic autoimmune diseases, disease diagnosis is often delayed. I am leveraging real-world patient data from a national consortium of medical institutions (eMERGE) to determine the predictive ability of polygenic risk scores (PRS) in stratifying severity of lupus and patients’ predisposition to manifesting other autoimmune conditions, rheumatoid arthritis and scleroderma. This study is novel in its simultaneous use of genetic and clinical patient data in augmenting early diagnosis, population health management screening, effective therapeutic management, and care for people with lupus.
Publications and Presentations
Tuteja, Sachleen, Sabah Kadri, and Kai Lee Yap. A performance evaluation study: Variant annotation tools-The enigma of clinical next generation sequencing (NGS) based genetic testing. Journal of Pathology Informatics 13 (2022): 100130. Available here.
Tuteja, Sachleen, Noah Forrest, and Theresa Walunas. Investigating the shared genetic underpinnings in systemic lupus erythematosus and rheumatoid arthritis. American Medical Informatics Association Annual Symposium, 2022. Available here.
Tuteja, Sachleen, Sabah Kadri, and Kai Lee Yap. An evaluation of variant annotation tools – Alamut Batch, ENSEMBL Variant Effect Predictor (VEP), and ANNOVAR – for clinical next generation sequencing (NGS) based genetic testing. American Biomedical Research Conference for Minoritized Scientists, April 26, 2022. Available here.
Tuteja, Sachleen, Sabah Kadri, and Kai Lee Yap. The enigma of clinical next generation sequencing based genetic testing – variant annotation tools: A performance evaluation study. American Medical Informatics Association Annual Symposium, 2021. Available here
Tuteja, Sachleen, Sabah Kadri, and Kai Lee Yap. The enigma of clinical next generation sequencing based genetic testing – variant annotation tools: A performance evaluation study. Japan Super Science Fair, 2021. Available here.